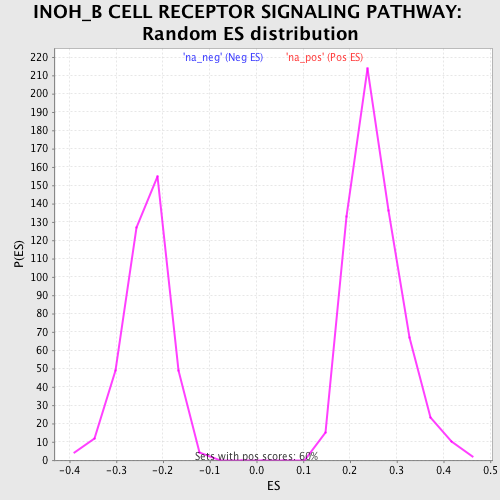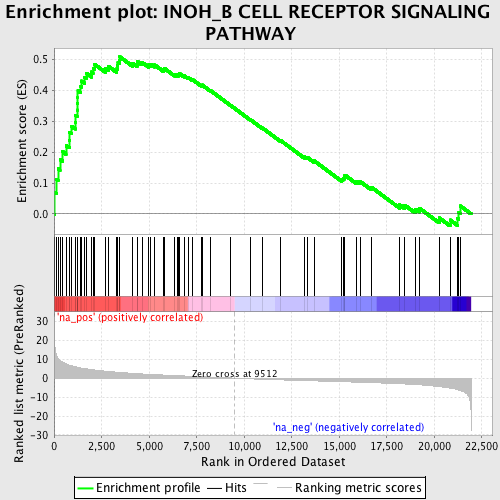
Profile of the Running ES Score & Positions of GeneSet Members on the Rank Ordered List
| Dataset | set02_BT_ATM_minus_versus_ATM_plus |
| Phenotype | NoPhenotypeAvailable |
| Upregulated in class | na_pos |
| GeneSet | INOH_B CELL RECEPTOR SIGNALING PATHWAY |
| Enrichment Score (ES) | 0.50804865 |
| Normalized Enrichment Score (NES) | 2.003886 |
| Nominal p-value | 0.0 |
| FDR q-value | 0.005759335 |
| FWER p-Value | 0.107 |

| PROBE | GENE SYMBOL | GENE_TITLE | RANK IN GENE LIST | RANK METRIC SCORE | RUNNING ES | CORE ENRICHMENT | |
|---|---|---|---|---|---|---|---|
| 1 | LCK | 20 | 19.234 | 0.0702 | Yes | ||
| 2 | NFKBIA | 116 | 12.486 | 0.1121 | Yes | ||
| 3 | PIK3CD | 211 | 10.436 | 0.1464 | Yes | ||
| 4 | TRAF3 | 312 | 9.412 | 0.1766 | Yes | ||
| 5 | TRAF4 | 456 | 8.404 | 0.2012 | Yes | ||
| 6 | VAV1 | 628 | 7.582 | 0.2214 | Yes | ||
| 7 | TRAF1 | 801 | 6.898 | 0.2391 | Yes | ||
| 8 | RAF1 | 819 | 6.824 | 0.2635 | Yes | ||
| 9 | RELA | 916 | 6.549 | 0.2834 | Yes | ||
| 10 | MAP2K2 | 1109 | 6.063 | 0.2970 | Yes | ||
| 11 | GAB1 | 1132 | 6.018 | 0.3183 | Yes | ||
| 12 | SYK | 1212 | 5.837 | 0.3362 | Yes | ||
| 13 | CBL | 1220 | 5.822 | 0.3575 | Yes | ||
| 14 | ARAF | 1249 | 5.744 | 0.3774 | Yes | ||
| 15 | TRAF2 | 1256 | 5.730 | 0.3983 | Yes | ||
| 16 | LCP2 | 1403 | 5.391 | 0.4116 | Yes | ||
| 17 | CD79B | 1433 | 5.349 | 0.4301 | Yes | ||
| 18 | GAB2 | 1612 | 5.026 | 0.4405 | Yes | ||
| 19 | DOK2 | 1695 | 4.902 | 0.4549 | Yes | ||
| 20 | MAPK3 | 1966 | 4.511 | 0.4593 | Yes | ||
| 21 | CRADD | 2050 | 4.400 | 0.4717 | Yes | ||
| 22 | ITPR1 | 2124 | 4.312 | 0.4843 | Yes | ||
| 23 | SHC1 | 2716 | 3.694 | 0.4710 | Yes | ||
| 24 | PIK3CA | 2870 | 3.541 | 0.4771 | Yes | ||
| 25 | DOK1 | 3306 | 3.173 | 0.4689 | Yes | ||
| 26 | IRS2 | 3324 | 3.163 | 0.4799 | Yes | ||
| 27 | SOS1 | 3360 | 3.127 | 0.4898 | Yes | ||
| 28 | FADD | 3443 | 3.070 | 0.4974 | Yes | ||
| 29 | BRAF | 3459 | 3.052 | 0.5080 | Yes | ||
| 30 | CRKL | 4137 | 2.560 | 0.4866 | No | ||
| 31 | BCAR1 | 4367 | 2.400 | 0.4850 | No | ||
| 32 | NFKBIB | 4385 | 2.393 | 0.4931 | No | ||
| 33 | SRC | 4632 | 2.239 | 0.4901 | No | ||
| 34 | SH2D2A | 4976 | 2.050 | 0.4820 | No | ||
| 35 | FRS2 | 5085 | 1.984 | 0.4844 | No | ||
| 36 | NFKBIE | 5291 | 1.879 | 0.4820 | No | ||
| 37 | PIK3R1 | 5770 | 1.654 | 0.4662 | No | ||
| 38 | GRAP2 | 5821 | 1.627 | 0.4700 | No | ||
| 39 | NRAS | 6347 | 1.384 | 0.4511 | No | ||
| 40 | PRKCQ | 6476 | 1.323 | 0.4501 | No | ||
| 41 | FYN | 6571 | 1.275 | 0.4506 | No | ||
| 42 | TRAF6 | 6611 | 1.253 | 0.4534 | No | ||
| 43 | PRKCD | 6849 | 1.149 | 0.4468 | No | ||
| 44 | GRB7 | 7077 | 1.049 | 0.4403 | No | ||
| 45 | PRKCH | 7283 | 0.959 | 0.4345 | No | ||
| 46 | ITPR3 | 7763 | 0.754 | 0.4154 | No | ||
| 47 | SOS2 | 7785 | 0.747 | 0.4172 | No | ||
| 48 | SH2B3 | 8222 | 0.568 | 0.3994 | No | ||
| 49 | LAT | 9292 | 0.110 | 0.3509 | No | ||
| 50 | MYD88 | 10348 | -0.356 | 0.3040 | No | ||
| 51 | CD19 | 10959 | -0.570 | 0.2782 | No | ||
| 52 | IRS4 | 11904 | -0.890 | 0.2384 | No | ||
| 53 | IRS1 | 13166 | -1.265 | 0.1854 | No | ||
| 54 | MAP2K1 | 13318 | -1.307 | 0.1834 | No | ||
| 55 | PRKCZ | 13684 | -1.423 | 0.1719 | No | ||
| 56 | CD79A | 15127 | -1.844 | 0.1128 | No | ||
| 57 | SHC2 | 15204 | -1.866 | 0.1163 | No | ||
| 58 | GRB10 | 15277 | -1.887 | 0.1200 | No | ||
| 59 | PRKCI | 15299 | -1.894 | 0.1260 | No | ||
| 60 | BLNK | 15915 | -2.099 | 0.1057 | No | ||
| 61 | PRKCE | 16103 | -2.152 | 0.1051 | No | ||
| 62 | FRS3 | 16687 | -2.351 | 0.0871 | No | ||
| 63 | HRAS | 18182 | -2.929 | 0.0297 | No | ||
| 64 | SHC3 | 18446 | -3.048 | 0.0289 | No | ||
| 65 | PIK3R2 | 19008 | -3.358 | 0.0157 | No | ||
| 66 | GRB14 | 19214 | -3.487 | 0.0192 | No | ||
| 67 | KRAS | 20270 | -4.421 | -0.0127 | No | ||
| 68 | GRB2 | 20842 | -5.251 | -0.0193 | No | ||
| 69 | NFKB1 | 21220 | -6.043 | -0.0142 | No | ||
| 70 | CRK | 21299 | -6.288 | 0.0055 | No | ||
| 71 | PRKCA | 21377 | -6.512 | 0.0261 | No |